Bacteria thrive on our planet in extreme conditions for billions of years. Their adaptive success depends on the ability to sense environmental cues and rapidly change their behavior. Two related small signaling molecules, (p)ppGpp and c-di-GMP, both discovered decades ago, are known to operate independently in regulating different aspects of bacterial growth, behavior, and stress response. Our new research now shows that these two global signaling molecules antagonize each other and that they can converge on a common target protein to control bacterial growth and metabolism converging 1. That the two signaling molecules interact could have been expected given their global regulatory roles in a wide range of bacteria and their similar chemical nature. Also, they were shown to inversely control cellular programs in Caulobacter crescentus, a riveting model organism with a characteristic bimodal life cycle. Whereas (p)ppGpp arrests Caulobacter in its free-swimming, non-reproductive state, c-di-GMP promotes the transition to a sessile and surface-attached mode of growth. Moreover, we had found that (p)ppGpp and c-di-GMP reciprocally control Caulobacter growth on glucose. However, the molecular mechanism of this interplay remained unknown.
The development of biochemical methods to identify effectors of small signaling molecules greatly complemented functional metagenomics uncovering a collection of novel proteins binding c-di-GMP or ppGpp 2-3. In our laboratory, putative c-di-GMP effectors were identified by directly capturing them from Caulobacter cell extracts. This, together with biochemical validation of candidates revealed several novel c-di-GMP binding proteins. One of these, a protein of unknown function that we named SmbA, showed homology to enzymes involved in redox reactions, pointing to a possible role in cellular homeostasis. Initial attempts to connect SmbA to some of the well-known c-di-GMP-dependent cellular processes such as biofilm formation, motility, or cell cycle control were not successful. However, the first step towards understanding the role of SmbA came from a simple control experiment, in which we test binding specificity and found that ppGpp, could effectively compete with c-di-GMP for SmbA binding. NMR titration experiments and crystal structures of SmbA with two different ligands occupying the same binding pocket left no doubt that we had identified a common target for two global second messengers (Figure 1).

Figure 1. Competitive binding of two second messengers to the same SmbA binding pocket.
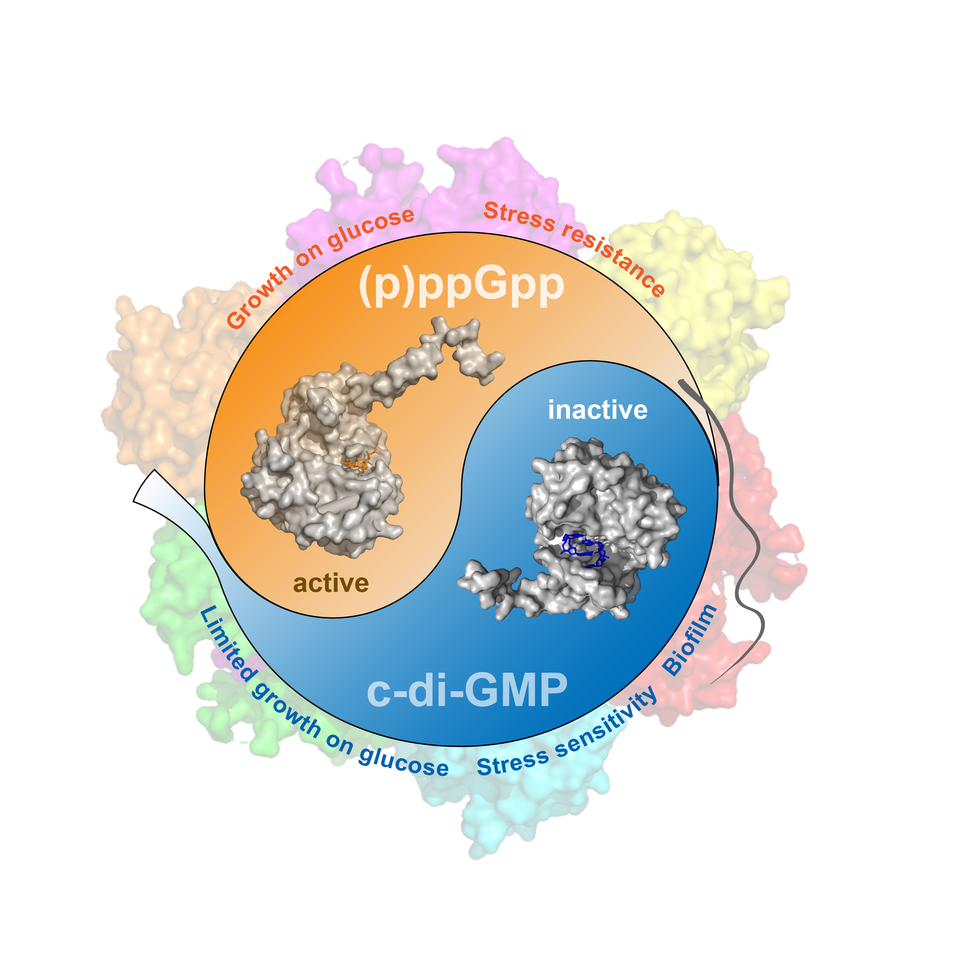
The concentrations of (p)ppGpp and c-di-GMP in the cell are influenced by nutritional conditions, stress factors, and other environmental cues. This information is read out directly by the SmbA protein, which can bind both molecules and acts as a molecular switch (Figure 2). We propose that in the non-dividing swimmers or in nutritionally stressed Caulobacter cells SmbA is switched on either because (p)ppGpp is high or because c-di-GMP is low. In this conformation, SmbA promotes glycolysis and neutralizes the production of harmful oxygen radicals, thereby maintaining the cells’ energy status and averting cell damage. In contrast, under conditions favoring growth and in dividing surface attached Caulobacter cells, c-di-GMP increases and displaces (p)ppGpp from the protein’s binding pocket. This leads to SmbA inactivation and to a rebalancing of carbon fluxes to promote the production of building blocks needed for Caulobacter growth and surface adherence.

Figure 2. Reciprocal control of SmbA by (p)ppGpp and c-di-GMP
mediating Caulobacter growth and development on glucose.
DGC, diguanylate cyclase, PDE, phosphodiesterase.
Caulobacter is a harmless organism that contributes to maintaining the metabolic cycles in the environment. However, both ppGpp and c-di-GMP are known to influence virulence, persistence, and antibiotic resistance of human pathogens. We believe that this work has opened up entry points into future studies addressing how global regulatory networks interact and how this affects growth and behavior of important pathogens.





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in