The paper in Nature Microbiology is here: https://rdcu.be/K2hN
This story begins
A long time ago in a seminar room far, far away. It’s where InfectX, a large multi-lab research project with the goal to comprehensively identify the components of the human infectome, was created. Here, a chance encounter between Bernd Wollscheid and myself initiated what is now a long-standing and productive collaboration.
Technology drives biology
In our first collaborative project we developed an innovative chemoproteomic approach to discover cell-surface receptors for known ligands. Using TRICEPS-based Ligand Receptor Capture (TRICEPS-LRC) technology to label intact mature vaccinia viruses, we identified the cell surface proteins AXL, M6PR, DAG1, CSPG4 and CDH13 as binding factors on human cells. In the context of discussions about this first project we became interested looking at the impact of vaccinia virus (VACV) infection on cell signaling. However, the complexity of cell signaling networks with built in redundancy, feedback loops and multi-enzyme/substrate relationships makes them difficult to dissect using the currently available toolboxes To truly unravel a signaling network, we needed a closed system containing a defined subset of players, and no redundancy.
VACV encodes two dual-specificity enzymes: F10 kinases and H1 phosphatase. F10 drives virus assembly and H1 assures virion transcriptional competence. Both enzymes are essential for infectious virus production (i.e. no redundancy), and both modify virus substrates (giving us a defined set of proteins to interrogate). With Karel Novy in Bernd’s lab taking the helm, we defined the vaccinia virus F10/H1 signaling network using a combination of proteotype analysis using quantitative mass-spectrometry and mutant F10(-) and H1(-) viruses. We found 33 F10/H1 phosphosites on 17 viral proteins. Excitingly, this including 8 phosphosites within 6 unknown shared substrates.
To cleave or not to cleave
Amongst these was the viral protease I7, the protein responsible for turning immature virions into mature virions through protein cleavage. Enter Samuel Kilcher from my lab. Four phospho-mutants, 3 cleavage assays, 2 transient complementations, and 1 Rif-release later and we know that F10 regulates I7 activity during virion maturation. Meanwhile Karel has mapped the peptide abundance of I7 substrates in wild-type and H1(-) virions, and given birth to the "Novy plot" (Figure 1), which shows that I7 substrates are hypercleaved in the absence of H1. Put simply, F10 phosphorylation turns on I7 protease activity and H1 de-phosphorylation shuts I7 down.
Figure 1: H1 regulates virion-associated I7 proteolytic activity. "Novy Plot" from Novy et al.. The relative change in abundance of tryptic peptides measured for structural proteins between H1(-) and WT MVs are displayed on the proteins’ linear sequence from N- to the C- terminus.
Lost in transcription
Something was missing. We identified 179 phosphosites in 47 viral proteins, yet none of them accounted for the transcriptional defect in H1(-) virions? Intriguingly, although H1 is a dual-specificity enzyme we didn’t identified a single tyrosine phosphosite. “Below detection and FYI, at least in eukaryotes protein phosphorylation commonly occurs on serine, threonine and tyrosine amino acids at an estimated ratio of 1000:100:1”, Karel said, “we need 5 milligrams of VACV for a quantitative phosphoproteomic profiling experiment and change the strategy a bit”. I had to laugh, it’s the only time I’ve ever weighed a virus prep. We found that 3 components of the viral transcription machinery were hyper-phosphorylated on tyrosines residues in H1(-) virions. Six phospho-mutants, 3 transient complementations, 1 new fluorescent recombinant virus, and a slew of EMs later, Samuel has shown that loss of dynamic phosphorylation of A7, the viral early transcription factor, underlies the H1(-) transcription defect.
From Zürich to Melbourne & London and back
In sum, these results establish a key role for dynamic phosphorylation in the regulation of infectious poxvirus particle assembly and highlight the utility of combining quantitative proteomic screens with mutant viruses to uncover proteotype-phenotype-genotype relationships. The last year of manuscript preparation and the revision was performed while Bernd was in Zürich, Karel was in Melbourne, and Samuel and I were in London. Karel and Samuel have since returned to Zürich, making this manuscript both a cross-disciplinary and a cross-continental effort to uncover a complex viral signaling network that controls virus assembly and transcriptional competence (Figure 2).
Figure 2: Model of poxvirus assembly driven by F10 kinase and H1 phosphatase. Summary figure from Novy et al.
The paper in Nature Microbiology is here: https://rdcu.be/K2hN
Written by Jason Mercer and Bernd Wollscheid on behalf of the other authors.
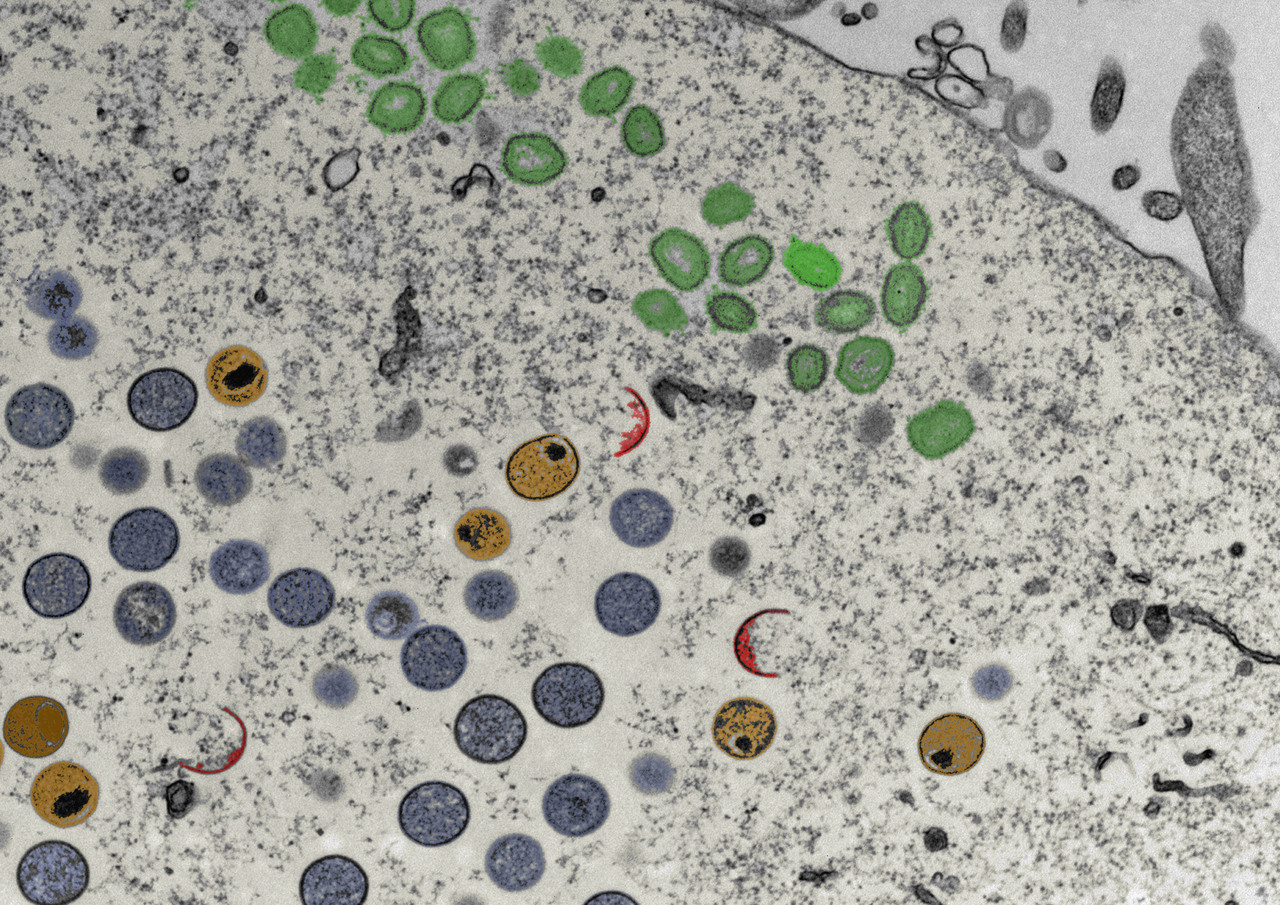
Poster EM image credit to Ian White; MRC-LMCB UCL: The red objects are called viral crescents, which grow to form the violet circles called immature virions, when the viral genome is encapsidated into these they become known as immature virions with nucleoid (the genome is the very dense structure within), the green objects are fully assembled infectious vaccinia viruses, which are called mature virions.
- Jason Mercer is a principal investigator & MRC group leader at the University College London, United Kingdom
- Bernd Wollscheid is a Professor at the Department for Health Sciences and Technology of ETH Zürich, Switzerland



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in